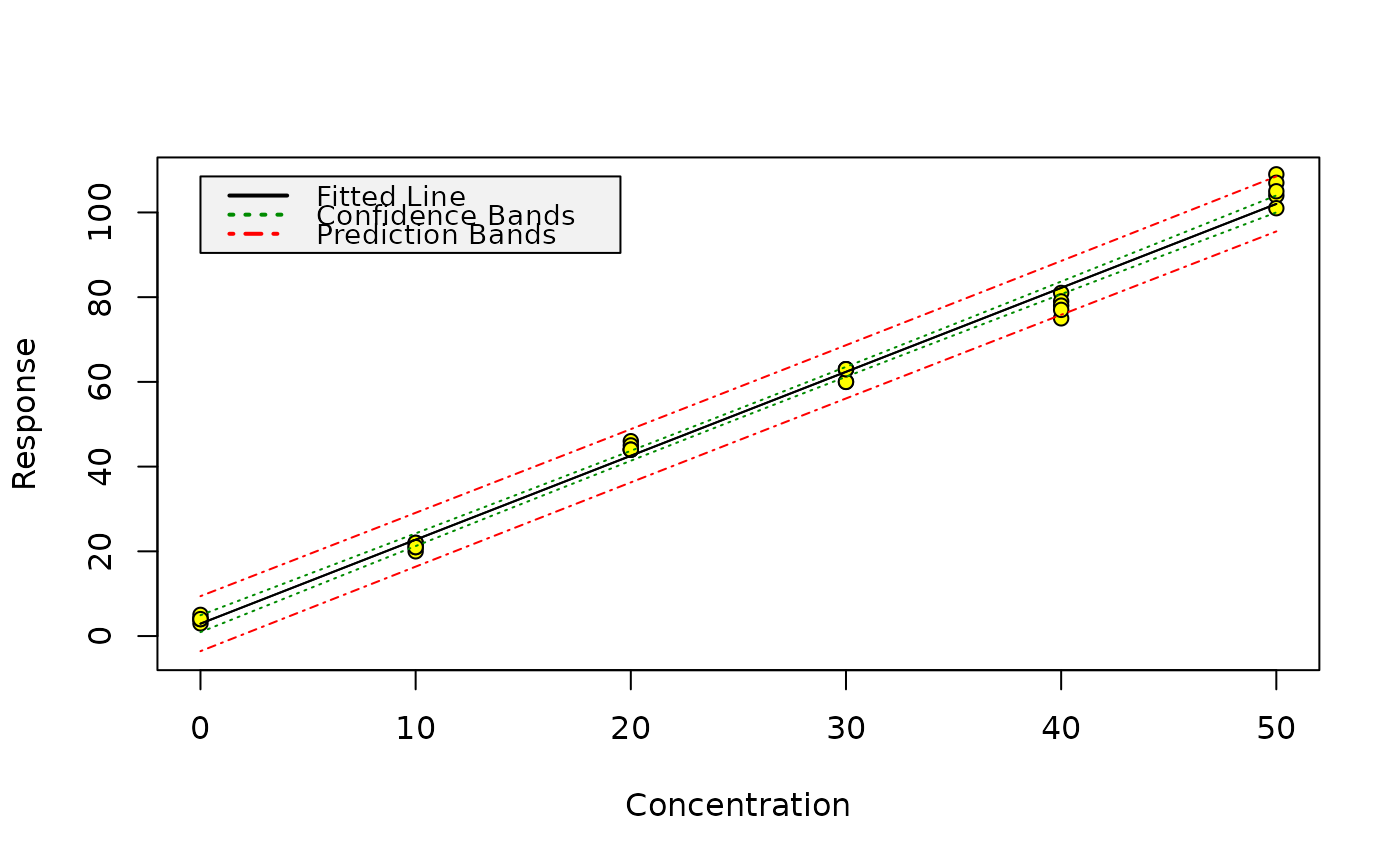
Produce graphics of calibration data, the fitted model as well as confidence, and, for unweighted regression, prediction bands.
Arguments
- object
A univariate model object of class
lmorrlmwith model formulay ~ xory ~ x - 1.- xlim
The limits of the plot on the x axis.
- ylim
The limits of the plot on the y axis.
- xlab
The label of the x axis.
- ylab
The label of the y axis.
- legend_x
An optional numeric value for adjusting the x coordinate of the legend.
- alpha
The error tolerance level for the confidence and prediction bands. Note that this includes both tails of the Gaussian distribution, unlike the alpha and beta parameters used in
lod(see note below).- varfunc
The variance function for generating the weights in the model. Currently, this argument is ignored (see note below).
Value
A plot of the calibration data, of your fitted model as well as lines showing the confidence limits. Prediction limits are only shown for models from unweighted regression.
Note
Prediction bands for models from weighted linear regression require
weights for the data, for which responses should be predicted. Prediction
intervals using weights e.g. from a variance function are currently not
supported by the internally used function predict.lm,
therefore, calplot does not draw prediction bands for such models.
It is possible to compare the calplot prediction bands with
the lod values if the lod() alpha and beta parameters
are half the value of the calplot() alpha parameter.